This is the first in a series of posts written by the engineers and developers of Wildbook. Drew has been with Wild Me since 2015, working mostly on Flukebook, the Wildbook for Cetaceans, and more recently doing cross-species algorithm-development as a Machine Learning Engineer.
We use a lot of different computer vision algorithms here at Wild Me. Some are old-school like the pattern-matcher used on Whaleshark.org, where users manually click the spots on a whale shark, and the Modified Groth algorithm does individual ID using nothing more than these spot coordinates and some trigonometry. More advanced is the widely-deployed HotSpotter algorithm, which can ID humpback whale flukes, zebras, cheetahs, and all sorts of animals that have distinct patterns. HotSpotter automatically extracts pattern-features from photos without any user input, then matches these features by analyzing the patterns of local pixel contrast within them. Even more advanced are our machine learning techniques, especially those using deep neural networks. This category includes our detector, which uses convolutional neural nets trained to draw bounding boxes around animals in photos, as well as neural net classifiers like the Deepsense algorithm that matches north atlantic right whales on Flukebook.
In this post, I’m introducing the newest, shiniest algorithm in our quiver of matching methods, one that marries the accuracy of neural networks with the flexible architecture of methods like HotSpotter. Developed by computer science PhD student Olga Moskvyak at Queensland University of Technology and productized by myself, we’re proud to add the PIE algorithm to Wildbook. PIE stands for “Pose Invariant Embeddings”, taken from the title of Moskvyak’s paper Robust Re-identification of Manta Rays from Natural Markings by Learning Pose Invariant Embeddings. We’ll be looking at how PIE works, where it’s applied so far, and some future avenues we’re exploring with this awesome new technique.
What is PIE?
The core concepts we need to explain PIE are embeddings and neural networks. Understanding these, we’ll see how PIE is a flexible and powerful tool for individual ID.
First, embeddings. In computer vision, an embedding is an abstract, numerical representation of an image that allows us to make semantic decisions based on that representation rather than the image itself. To explain through example, the spot-matcher on Whaleshark.org works on spot-coordinates; these coordinates constitute the embedding for a given image and the Modified Groth algorithm matches these embeddings using geometry. All of this ultimately allows us to ask “which animal is this?”.
Now for neural networks. Neural networks are the most popular technique in modern machine learning. At a high level, they’re inspired by how neurons work in brains, able to learn and perform all sorts of tasks by adjusting a huge network of small, flexible units. Practically speaking, neural networks are systems that can be iteratively improved to perform a task on training data. This process is what we call training, or learning. For example the Deepsense right whale matcher on Flukebook is a neural network that was trained to look at an image of a right whale and return that whale’s ID according to the NARW catalog. When the network first started training it was very poor at that task, but through the training process it gradually improved to be quite accurate, at which point we made it available to all of you.
Unlike Deepsense, the PIE neural network is not trained to classify images into bins (one bin for each whale in the catalog). Instead, its deep neural network is trained to extract embeddings from images. Give an image to PIE, and it returns a list of 256 numbers between 0 and 1 (also called a 256-long or 256-dimensional vector): this is the image’s embedding. How does it choose those 256 numbers? That’s the trick, both clever and intuitive once you get it.
We call the 256-dimensional space of all possible PIE embeddings, fittingly enough, the embedding space. So PIE generates a mapping from an input image to a point in embedding space. Now, how are these embeddings chosen? How is PIE trained? Remember, the goal here is individual ID. PIE is trained with the simple concept that images of the same individual should produce similar embeddings, and images of different individuals should make different embeddings; the distance between two images in embedding space corresponds to the similarity between those images for the purpose of individual ID. The result of this is that an individual with N photos in the catalog should produce N points in embedding space all clustered close together, in a cluster unique to that individual. A second individual with M photos should be a distinct cluster elsewhere in embedding space. These two criteria together (same individual->close embeddings; different individuals->far) are the training signal for PIE, the metric that’s procedurally optimized during learning. The technical term for this measure is Triplet Loss, if you want to read more about that.
PIE is trained to learn embeddings that are useful for individual ID. Unlike HotSpotter, which is a “static” pattern matcher, ie a fixed algorithm not trained for each separate species (which by the way, is very impressive considering how broadly-effective HotSpotter is!), PIE can be trained on a per-species basis. So we have a separate PIE model optimized for manta rays versus humpback whales. And unlike fixed-catalog classifiers like Deepsense, PIE can gracefully add new individuals to its catalog without being retrained: it learns the general task of mapping images into embeddings that represent individuals, rather than the specific task of sorting images into a fixed number of IDs. PIE strikes a lovely balance between a flexible general-purpose identifier and one that can be trained and refined on a given problem.
How did PIE get into Wildbook?
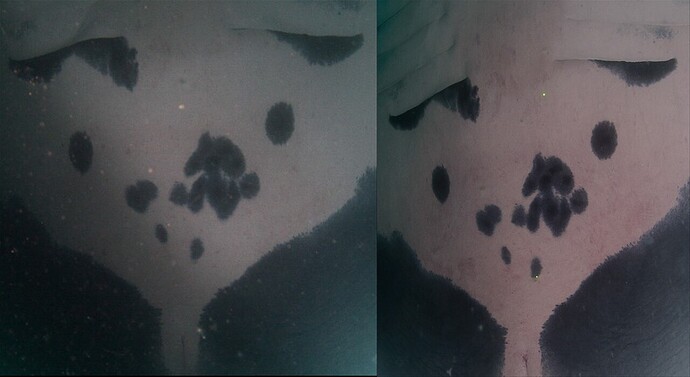
As you may have guessed from the title of Olga’s paper, PIE was developed with manta ray identification in mind. Manta ray bellies have individually-distinct patterning, but these patterns have proven challenging for HotSpotter and other techniques to match accurately in the past. This not only makes mantas a great subject for new research, but means that mantamatcher.org (the Wildbook for–you guessed it–manta rays (and other rays!)) the target of our first PIE deployment.
Moskvyak’s publicly-available PIE model was trained on a high quality curated dataset from Project Manta out of University of Queensland, who are themselves contributors to MantaMatcher, so we simply deployed that model on MantaMatcher.org. It’s still too early for us to have usage-based accuracy statistics, but the reported accuracy is 62% top-1 and 97% top-10 (meaning the algorithm returns the correct match in the first 10 candidates 97% of the time; as with all our ID algorithms, PIE return a list of candidate matches and relies on researchers to confirm the accuracy of those results). After years of being a Wildbook that doesn’t use advanced computer vision, MantaMatcher was upgraded to the absolute cutting edge of animal individual ID.
We take pride in being open source, and we are grateful that Olga Moskvyak shares open source values. Not only are the models from her publication publicly available, but more importantly the code she used to train those models. This crucially allows us to extend PIE to new species and train it on new datasets. We at Wild Me have access to some of the highest-quality animal ID data in the world, so this is a dream come true. The original stand-alone python program constituting PIE is available on github, so we forked that code into our wildbook-ia repository where all of our computer vision code lives. My task as a machine learning engineer these past few months has been to integrate the original PIE code into our wildbook-ia server so that it operates as a seamless component of the larger Wildbook platform. As mentioned above, this was implemented first on the MantaMatcher platform. And with that deployment complete, I’ve spent all my time since then applying PIE to new species.
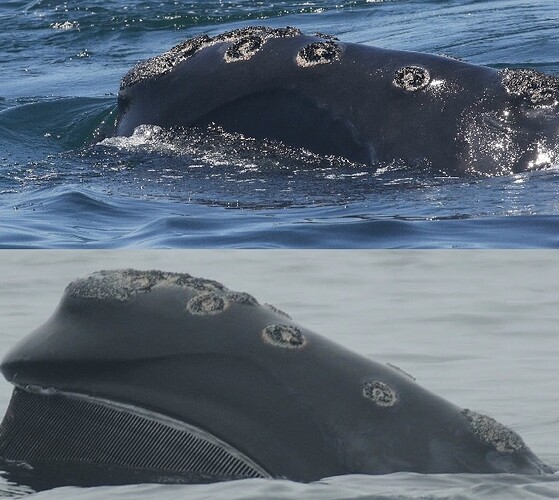
A researcher-confirmed match on a right whale lateral head callosity made by PIE.
Where else is PIE used?
Our first novel application of PIE was for lateral photos of right whale heads. As mentioned previously, there’s already a highly-accurate model for matching right whales on Flukebook, the Deepsense classifier. This model was trained on aerial photos, taken from aircraft or drone, that show clear and consistent views of the callosities on a right whale’s head. However, researchers don’t always have access to aircraft, and boat-based photos are different enough from aerial photos that the models are not cross-applicable. So there was a need from our right whale researchers to match the callosities on a whale’s head based on lateral, boat-based photos. With funding from NOAA on our ongoing right whale-matching collaboration, I trained and tuned the PIE architecture to the lateral right whale problem.
As is almost always the case with machine learning, there are two aspects of making a new model: there’s the obvious one, finding a new set of training data and re-training the system on that, but there’s also the less obvious problem of finding the right big-picture settings, called “metaparameters” or “hyperparameters”, that define how exactly the training system works. For example, the learning rate of a neural network defines how much the network changes at each step of the training process. If your learning rate is too high, you might not get a very accurate solution because the network isn’t fine-tuned enough. But if it’s too low, training could take way too long: it could be the difference between a 4 hour training process and a 40 hour one. Another piece of the PIE architecture that we experimented with is called image augmentation, which is when you manipulate images in the training set to artificially increase the number of training images, for example stretching a photo, changing its exposure, or rotating it a few degrees. Image augmentation can prevent overfitting (when the network simply memorizes its training data and can’t extrapolate its reasoning to new images) so it produces more robust models, but if you use too much augmentation you might make the training data so complicated that the network just gets confused and never learns very well. The model-development process involves experimenting with these types of settings to make the most accurate system possible.
After a few weeks of working on the right whale lateral model, we arrived at a model with a top-12 accuracy of 90%. While not as accurate as say, the manta ray or humpback fluke matchers that approach 99% in this metric, we consider this a much more difficult problem considering the nature of these whale callosities and the photos in which they appear. This difficulty is also reflected in the fact that, to our knowledge, this is the first ever automatic system for matching boat-based photos of right whales. This new flavor of PIE was deployed on Flukebook at the beginning of October and is available to our users.
Later this week we’ll be deploying yet another novel species for PIE, one that’s even more challenging than lateral right whales: orcas! I grew up in the Pacifict Northwest of the United States, a place where we’re very proud of our resident (and transient!) killer whales, so this is super exciting to me. We started the effort with a dataset contributed by the Norwegian Orca Survey that we hand-annotated to draw bounding boxes around the animals. We had already curated this data as part of our development of the finFindR dorsal fin trailing-edge-matching algorithm, which is super accurate for bottlenose and other smaller dolphins, but has not been as impressive on orcas. Simply put, orca dorsal fin edges are not as distinct as bottlenose dolphins’, at least among the data we’ve seen. From conversations with the researchers we know in the orca community, we learned that the saddle patch just behind an orca’s dorsal fin is the most distinct and easily-photographed feature on the animals, so that became our target for matching with the PIE pattern-matcher. Unlike the pigment on a humpback whale’s fluke, these saddle patches are not dramatically different in terms of large shapes and patterns; instead, scars and other subtle features on the area are the distinguishing characteristics in these photos.
We found orcas to be much more challenging! Our latest PIE model achieved 60% top-12 accuracy on these charismatic cetaceans. However, as many researchers are well aware, manually matching field photos without algorithmic assistance can be an enormous time-sink, and making that job easier by any amount can be a big help. We are deploying this model on Flukebook as a time-saving tool for orca researchers, and alongside the finFindR trailing-edge matcher, that community will have access to two cutting-edge algorithms to assist their data curation efforts.
One technique we tried to improve orca accuracy was automatically removing background sea and sky from images.
As we’ve seen with orcas, no one tool is a magical solution that will ID every species perfectly well–not even PIE. But with open source software and collaboration with the field researchers who know these animals better than anyone, we can turn cutting-edge machine learning research into tools that help the field biologist study earth’s creatures. This is far from the end of our new developments with Olga Moskvyak’s algorithm: now that we’ve explored some new computer vision problems like orca saddle patches, we’re going to revisit some older problems such as whale sharks or aerial photos of right whales, to see if the latest deep learning techniques might improve the accuracy or workflow of identifying individual animals on Wildbook.
Addendum 8 December
With the help of some orca researchers, we found a pervasive data error in the orca sightings on Flukebook that were used to train PIE: a significant minority of our orca encounters had been mislabeled with the wrong individual. As you can imagine, machine learning models have a more difficult time when the ground truth training data is inaccurate, so this explains some of the lower performance we had seen matching these animals. After correcting this error, we found PIE to be much more effective on killer whales than first thought: top-12 validation accuracy rose to 84% vs the previous 60%; a huge improvement. This latest model is available now on Flukebook.